A Mining Software Repository and Ticket management Platform tool, in Pharo
# GitProject health
[](https://github.com/moosetechnology/GitProjectHealth/actions/workflows/ci-moose11.yml)
[](https://github.com/moosetechnology/GitProjectHealth/actions/workflows/ci-moose12.yml)
[](https://coveralls.io/github/moosetechnology/GitProjectHealth?branch=main)
[](https://doi.org/10.5281/zenodo.13886504)
This project includes a model, an importer, and some visualizations to evaluate the health of a GitLab or GitHub group.
## Installation
Download a [Moose image](https://modularmoose.org/beginners/install-moose/).
In the Moose image, in a playground (`Ctrl+O`, `Ctrl+W`), perform:
```st
Metacello new
repository: 'github://moosetechnology/GitProjectHealth:main/src';
baseline: 'GitLabHealth';
onConflict: [ :ex | ex useLoaded ];
onUpgrade: [ :ex | ex useIncoming ];
onDowngrade: [ :ex | ex useLoaded ];
load
```
## Usages
### Import
#### Group import: GitLab
In a playground (`Ctrl+O`, `Ctrl+W`):
```st
glhModel := GLHModel new.
repoApi := GitlabApi new
privateToken: '<my private token>';
hostUrl: 'https://gitlab.myPrivateHost.com/api/v4'.
gitlabImporter := GitlabModelImporter new
repoApi: repoApi;
glhModel: glhModel.
"137 is the ID of the a Group, you can find the number in the webpage of every project and group"
gitlabImporter importGroup: 137.
```
#### Group import: GitHub
In a playground (`Ctrl+O`, `Ctrl+W`).
```st
glhModel := GLHModel new.
githubImporter := GithubModelImporter new
privateToken: '<my private token>';
glhModel: glhModel.
githubImporter importGroup: 'moosetechnology'.
```
#### Extract more commits
> [!NOTE]
> GitLab API only.
You might want to gather more commits for a specific repository.
To do so for GitLab, we added the following API:
```st
myProject := (glhModel allWithType: GLHProject) detect: [ :project | project name = '<my project name>' ].
gitlabImporter importCommitsOfProject: myProject since: nil until: '2023-01-01'.
```
### Visualize
To visualize the group "health":
```st
myGroup := (glhModel allWithType: GLHGroup) detect: [ :group | group id = 137 ].
canvas := GLHGroupVisualization new forGroup: { myGroup }.
canvas open.
```
### Export
To export the visualization as an svg image:
```st
canvas svgExporter
withoutFixedShapes;
fileName: 'drit-group-health';
export.
```
## Metamodel
Here is the metamodel used in this project:
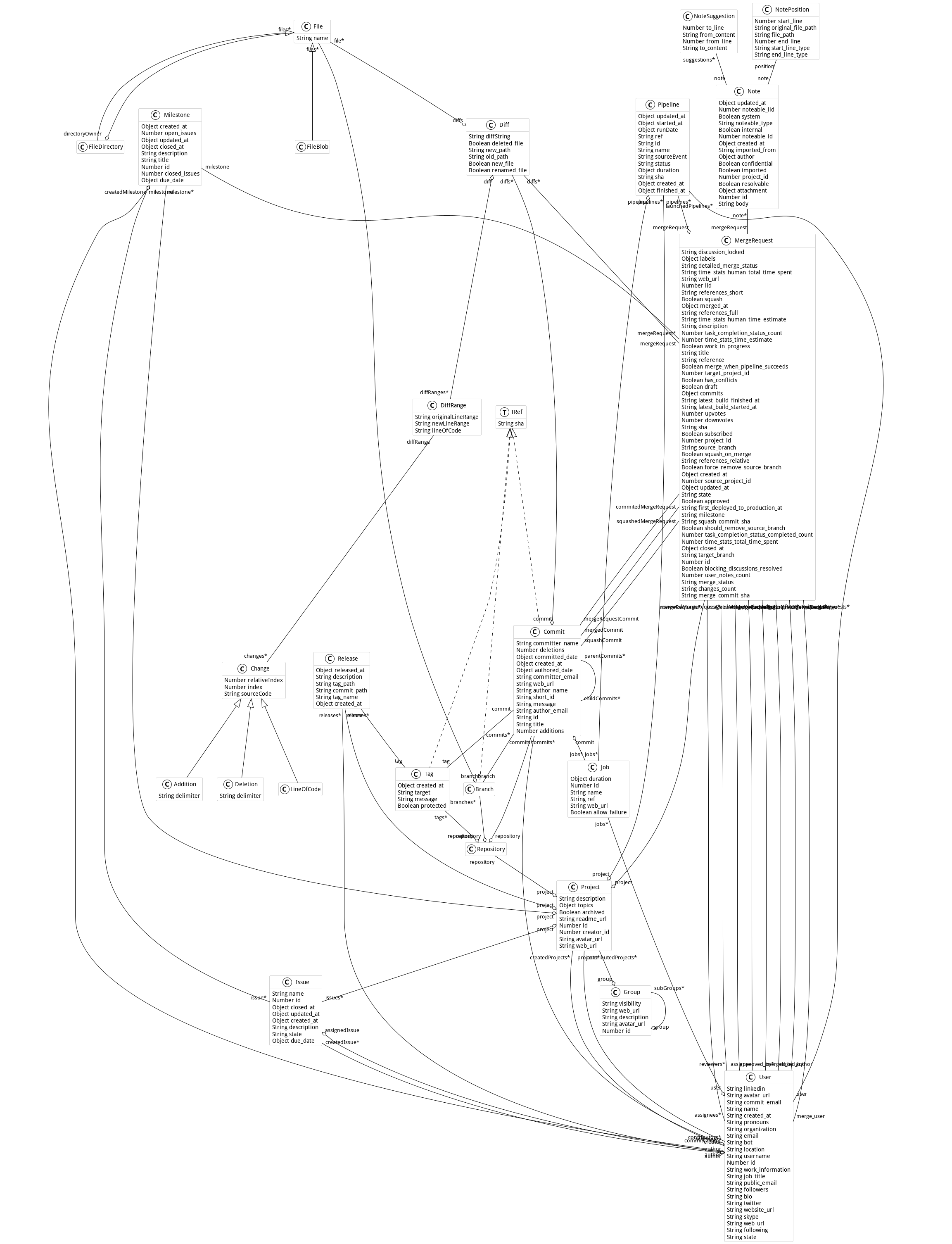
## Connectors
This project comes with connectors to other metamodels to increase its powerfulness.
Explore this part of the [documentation on the main website](https://modularmoose.org/users/git-project-health/getting-started-with-gitproject-health).
## Contributor
This work was first developed by the [research department of Berger-Levrault](https://www.research-bl.com/).
## Running metrics with docker
### Running locally
This example shows how to set up and run GitProjectHealth metrics for a given set of users and their projects over two periods of time.
It ouputs a csv files containing: code churn, commit frequencies, code additions and deletions, comments added (e.g., `//`, `#`, `/**/`), average delay before first churn, and merge request duration.
```smalltalk
| glhModel gitlabApi glhImporter |
"load GitProjectHealth into your image"
Metacello new
repository: 'github://moosetechnology/GitProjectHealth:main/src';
baseline: 'GitLabHealth';
onConflict: [ :ex | ex useIncoming ];
onUpgrade: [ :ex | ex useIncoming ];
onDowngrade: [ :ex | ex useLoaded ];
ignoreImage;
load.
"set up a log at your root"
TinyLogger default addFileLoggerNamed: 'pharo-code-churn.log'.
"new model instance"
glhModel := GLHModel new.
"new API class instance"
gitlabApi := GitlabApi new
privateToken: '<YOUR_TOKEN_KEY>';
hostUrl: 'https://gitlab.com/api/v4'.
"new importer instance"
glhImporter := GitlabModelImporter new
repoApi: gitlabApi;
glhModel: glhModel;
withFiles: false;
withCommitDiffs: true.
"export metrics, the output files are located at `FileLocator home / *.csv`"
GitMetricExporter new
glhImporter: glhImporter;
addAPeriodFrom: '1 march 2023' to: '24 may 2023';
addAPeriodFrom: '1 march 2024' to: '24 may 2024';
setupAnalysisForUsersWithNames: { 'John Doe' };
setupAnalysisForProjectsWithIds: { 14. 543. 2455 };
label: 'GitLabHealth';
exportInCSV.
```
### deploying with docker
```bash
git clone https://github.com/moosetechnology/GitProjectHealth.git
cd GitProjectHealth
sudo docker build -t code-churn-pharo .
sudo docker run code-churn-pharo &
```
Locate and retrieve csv output files:
```bash
sudo docker ps
sudo docker exec -it <container-id> find / -type f -name '*.csv' 2>/dev/null
```